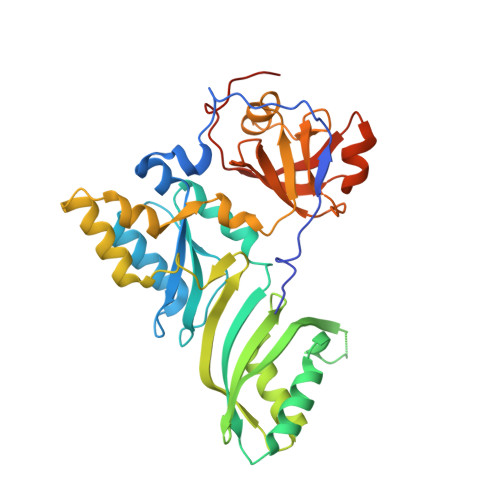
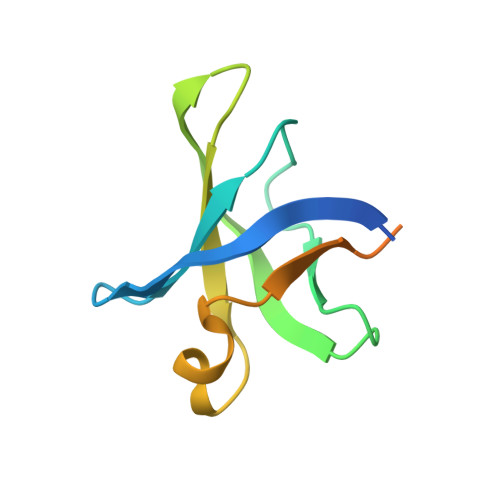
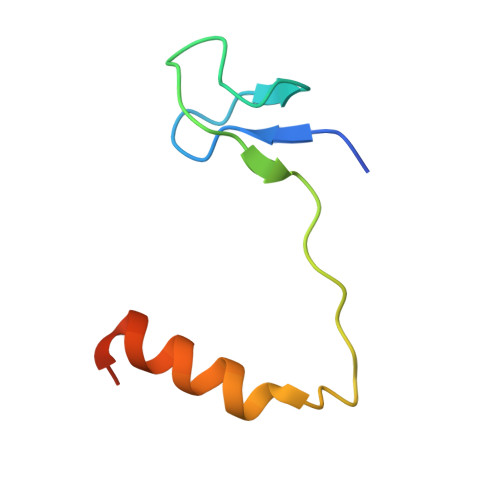
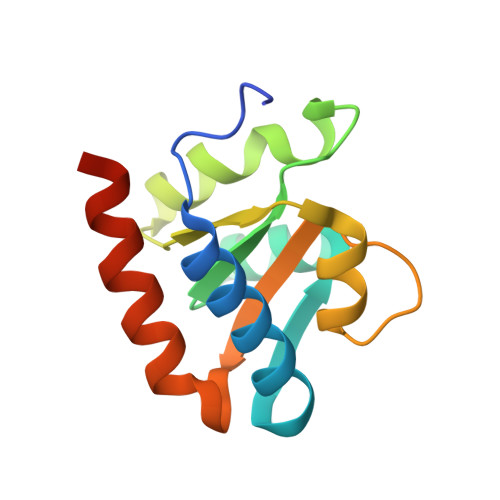
Crystal structure of an H/ACA box ribonucleoprotein particle
Li, L., Ye, K.(2006) Nature 443: 302-307
- PubMed: 16943774
- DOI: https://doi.org/10.1038/nature05151
- Primary Citation of Related Structures:
2HVY - PubMed Abstract:
H/ACA ribonucleoprotein particles (RNPs) are a family of RNA pseudouridine synthases that specify modification sites through guide RNAs. They also participate in eukaryotic ribosomal RNA processing and are a component of vertebrate telomerases. Here we report the crystal structure, at 2.3 A resolution, of an entire archaeal H/ACA RNP consisting of proteins Cbf5, Nop10, Gar1 and L7ae, and a single-hairpin H/ACA RNA, revealing a modular organization of the complex. The RNA upper stem is bound to a composite surface formed by L7ae, Nop10 and Cbf5, and the RNA lower stem and ACA signature motif are bound to the PUA domain of Cbf5, thereby positioning middle guide sequences so that they are primed to pair with substrate RNA. Furthermore, Gar1 may regulate substrate loading and release. The structure rationalizes the consensus structure of H/ACA RNAs, suggests a functional role of each protein, and provides a framework for understanding the mechanism of RNA-guided pseudouridylation, as well as various cellular functions of H/ACA RNP.
Organizational Affiliation:
National Institute of Biological Sciences, 7 Science Park Road, Zhongguancun Life Science Park, Beijing 102206, China.