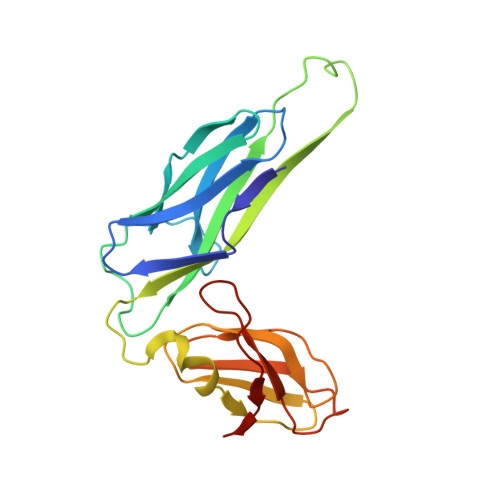
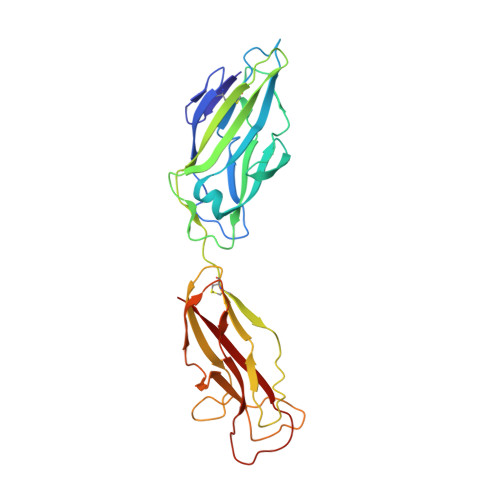
Structural basis of tropism of Escherichia coli to the bladder during urinary tract infection.
Hung, C.S., Bouckaert, J., Hung, D., Pinkner, J., Widberg, C., DeFusco, A., Auguste, C.G., Strouse, R., Langermann, S., Waksman, G., Hultgren, S.J.(2002) Mol Microbiol 44: 903-915
- PubMed: 12010488
- DOI: https://doi.org/10.1046/j.1365-2958.2002.02915.x
- Primary Citation of Related Structures:
1KIU, 1KLF - PubMed Abstract:
The first step in the colonization of the human urinary tract by pathogenic Escherichia coli is the mannose-sensitive binding of FimH, the adhesin present at the tip of type 1 pili, to the bladder epithelium. We elucidated crystallographically the interactions of FimH with D-mannose. The unique site binding pocket occupied by D-mannose was probed using site-directed mutagenesis. All but one of the mutants examined had greatly diminished mannose-binding activity and had also lost the ability to bind human bladder cells. The binding activity of the mono-saccharide D-mannose was delineated from this of mannotriose (Man(alpha 1-3)[Man(alpha 1-6)]Man) by generating mutants that abolished D-mannose binding but retained mannotriose binding activity. Our structure/function analysis demonstrated that the binding of the monosaccharide alpha-D-mannose is the primary bladder cell receptor for uropathogenic E. coli and that this event requires a highly conserved FimH binding pocket. The residues in the FimH mannose-binding pocket were sequenced and found to be invariant in over 200 uropathogenic strains of E. coli. Only enterohaemorrhagic E. coli (EHEC) possess a sequence variation within the mannose-binding pocket of FimH, suggesting a naturally occurring mechanism of attenuation in EHEC bacteria that would prevent them from being targeted to the urinary tract.
Organizational Affiliation:
Department of Molecular Microbiology, Washington University School of Medicine, St. Louis, MO 63110, USA.